|
7/30/2023 0 Comments Dfind old genome assemblies 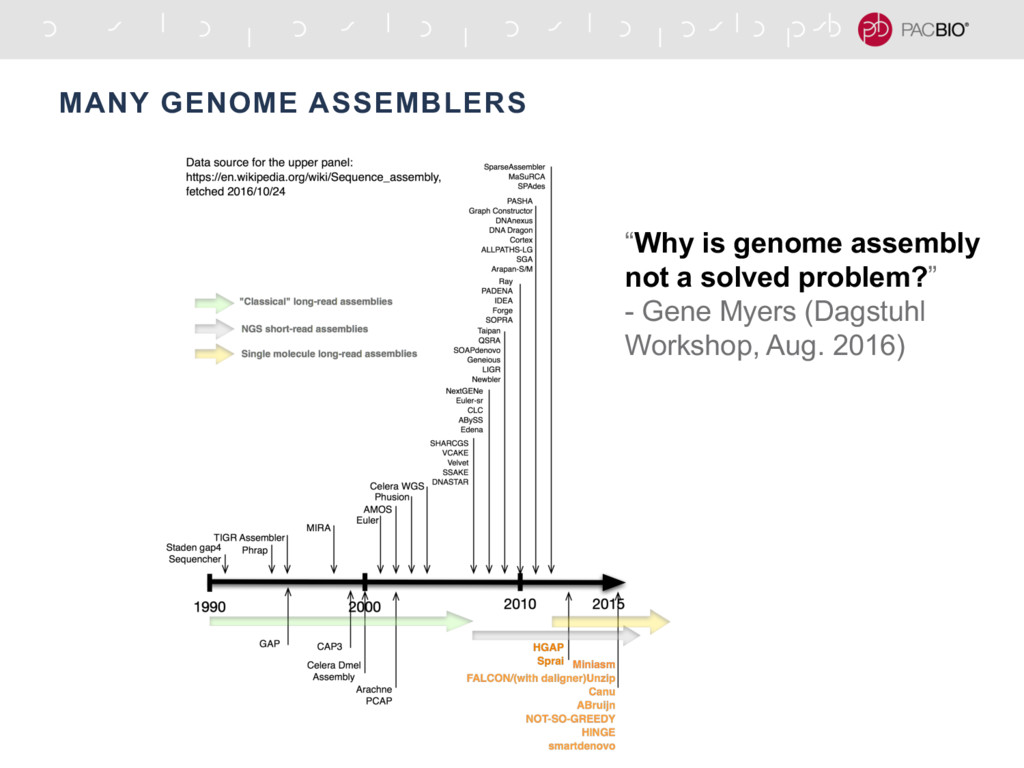 
Update to existing genome assembly ComponentĪt least one layer mandatory, with highest layer no lower than for existing assembly Tables I and II shows requirements for new genome assembly submissions and updates to existing assembly submissions. BAM alignment of reads to new chromosomeīrief information relating to assembly and future plansĬonsistent with the variety of assembly processes, submitters to INSDC approach with data for the layers in different combinations of layers. New genome assembly submissions ComponentĮ.g. The figure shows three typical assembly processes and theĪ) Clone-based assembly with scaffolding and finishing steps.ī) Shotgun assembly direct to chromosomes.Ĭ) Partial assembly to contigs only. Three typical assembly processesįigure I. This document lays out the requirements for submission of genome assembly information into INSDC databases. Genome assemblies comprise a number of possible layers of information, including reads, contigs, scaffolds and chromosomes (see figure I). The original site INSDC standards for genome assembly submission 2013.08.14 version INSDC standards for genome assembly submission INSDC standards for genome assembly submission.Use the BLAST search the assembly link under the Access the data section on the right side of the record to load the appropriate blastn suite.Access the record that you want, for example Algmis_Hirise_1.0.Search the Assembly database for your organism (for example, American Alligator).Unless you change the option, your search will be limited to your organism.įinally, there can be more than one assembly for the organism and you want to select your own. What happens if there is no genome assembly for the organism of your interest? You will learn that by using the BLAST Genomes search box when the software presents you with a blastn suite for searching the nucleotide collection (nt) instead. The Search Set Database menu is displaying the databases associated with the selected genome assembly If you have options, select the database to search or switch to the blastp suite for protein searches (by switching from the blastn tab to the blastp tab at the top of the page).įigure 2: Genome-specific blastn page. In this example the query sequence is NM_000249. To compare your sequence(s) against the genome, proceed as you would for any other blastn search, by entering your query sequence(s) in the search box as accession numbers or in the FASTA format. Moreover, there will also be genome-specific blastp suite for the proteins annotated on the assembly. In this example, there are 7094 sequences (emphasized with a red rectangle in Figure 2) that comprise the American Alligator genome assembly, ASM28112v4 . For NCBI-annotated assemblies, such as ASM28112v4, you will find additional databases of various transcript data. For assemblies that are not annotated, you will find a single database of sequences that comprise the assembly. The Search Set Database menu will display the associated databases. Upon your search, the BLAST software will display the genome-specific blastn suite (Figure 2) with the title reflecting the organism name. Select the matching name (taxid) for the organism from the listing, for example "American Alligator (taxid: 8496)"įigure 1: A part of the BLAST ® home page that includes the BLAST Genomes search box.Begin typing the name of the organism (for example "American") to invoke the organism listing.The BLAST Genomes search box, located on the BLAST ® (Basic Local Alignment Search Tool) home page (Figure 1) allows you to access the most complete* genome assemblies for eukaryotic organisms. Use these steps to locate such assemblies: 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
